Novel bacteria traps, heavy molecular labels, and UV chambers will be deployed by marine scientists at Davis this summer, to understand the sulphur-mediated attraction between Antarctic phytoplankton and bacteria.
The distinctive ‘smell of the sea’ is the result of an important sulphur-cycling relationship between phytoplankton and bacteria. But the intimate details of this liaison are not well known.
To find out more, marine biologist Dr Katherina Petrou and her colleagues from the University of Technology Sydney, are conducting a project* at Davis this summer to identify the bacteria involved and the effects of climate-related stressors on the sulphur cycle.
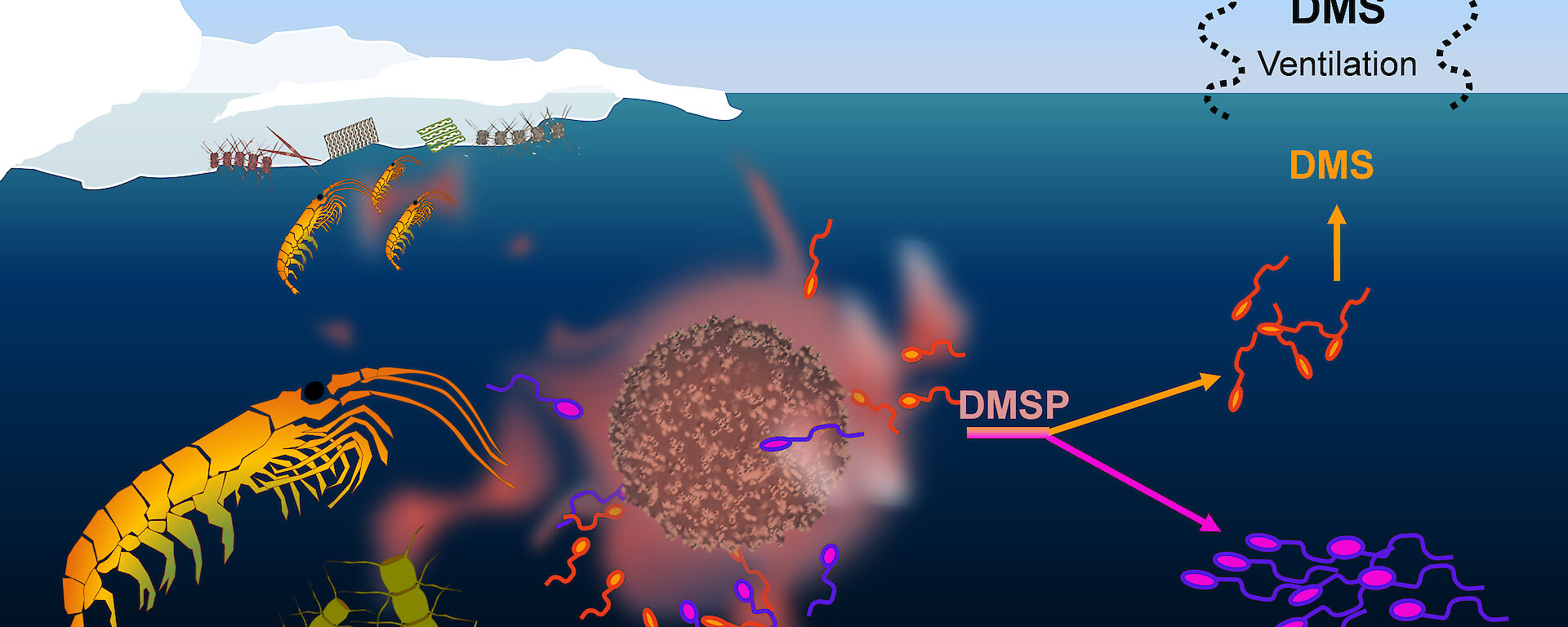
“We know that many species of phytoplankton, including Antarctic diatoms and haptophytes, transform sulphur in seawater into dimethylsulphoniopropionate (DMSP), which is very attractive to some bacterial species as a carbon and sulphur source,” Dr Petrou said.
“But we don’t know which bacteria are involved or how changes in DMSP production as a result of environmental stressors might influence the structure of the microbial community and subsequent chemical processes.”
This is important because when bacteria consume DMSP, released when phytoplankton die, they can either use it for their own growth needs, or metabolise it into dimethylsulphide (DMS). This volatile compound ventilates into the atmosphere, creating the sea smell reminiscent of boiled cabbage and forming aerosols that can promote cloud formation and subsequently influence regional climate.
One of the key questions Dr Petrou’s team wants to investigate is how increased ultraviolet light exposure affects phytoplankton production of DMSP and its subsequent fate. Phytoplankton generally exist in the sunlit layers of the ocean, from the surface down to about 100m depth. But as the ocean warms in response to climate change it can become more stratified, with warmer waters on top, trapping the phytoplankton in the uppermost, more exposed surface layer.
“One of the effects of ocean warming is that the surface mixed layer where phytoplankton reside becomes shallower, exposing the cells to more light and UV in a daily cycle,” Dr Petrou said.
“Studies elsewhere suggest that DMSP production increases under UV stress and we want to see whether this holds true in Antarctica and whether this shift affects the bacterial community and ultimately the fate of sulphur in the ocean.
“Will more DMS be released into the atmosphere or will the DMSP be utilised in the marine food web?”
To find out, Dr Petrou and her team will collect surface water samples (5–20m depth) at different locations and times during the five-week visit to Davis, to ensure they have a representative sample of the summer phytoplankton and bacterial community.
Some of these water samples will then be incubated in three-litre UV-transparent acrylic chambers, and exposed to different durations of UV light for up to three days. At the end of the experiment the amount of DMSP, and the numbers of phytoplankton and bacteria (including who is there), will be measured.
To identify the bacteria involved in the sulphur cycle, some of the water samples will be inoculated with DMSP tagged with a heavy carbon label (13C). When the bacteria consume this labelled DMSP some of the 13C will be incorporated into their DNA, making it heavier.
“We’ll be able to separate the heavier bacteria that have taken up the labelled DMSP from the lighter bacteria and then identify them through DNA sequencing,” Dr Petrou said.
The team will also use a novel micro-trap to capture DMSP-consuming bacteria in the water column. The trap relies on the principle of ‘chemotaxis’, where an organism moves in response to a chemical cue — in this case the attraction of bacteria to a food source.
“It’s a bit like a microbial mouse trap,” Dr Petrou said.
“We have some very tiny chambers containing DMSP, which we’ll deploy at different depths in the water column for about one hour. This will allow the DMSP to leak out into the surrounding water and form a chemical gradient that will be sensed by nearby bacteria. If they like what they sense they will swim down the gradient and into our chambers.”
The team will use DNA sequencing to identify the bacterial species captured, and they’ll be able to compare the identity of these captured bacteria with those from the 13C experiment, to see if they match.
“Marine microbes perform invaluable ecosystem services,” Dr Petrou said.
“Any changes in chemical cycling and the microbes responsible for that cycling will have ripple effects for the marine food web, affecting ocean productivity — including fisheries yields — and our climate.
“This project will provide the first insight into which direction the sulphur cycle might move in the future.”
Wendy Pyper
Australian Antarctic Division
*Australian Antarctic Science project 4315