Adélie penguins in Antarctica are an ideal species on which to study the speed and mechanisms of evolution. Recent research using modern and ancient samples of the birds’ DNA has shown that mutation rates in the DNA can be used as a proxy for the rate of evolution. The discovery could improve the dating of evolutionary events, such as the divergence of species from a common ancestor.
Studying evolution is not always easy. You need a lot of time and the opportunity to study many individuals. Also, it’s preferable to work on a species that has been unaffected by humans and that has lived in one place for a long time, in an environment that has changed a lot. If you're interested in ancient DNA, you’d prefer them to be ‘stored’ in cold conditions. Adélie penguins in Antarctica are ideal!
To most people evolution is the idea that the composition of the earth’s biodiversity has changed over long periods of time. To biologists it also means something more specific; namely changes in populations or species over shorter periods of time. At a practical level it usually comes down to changes in the ‘frequency’ (occurrence) of particular genes or DNA sequences over time.
How, precisely, might we go about detecting evolutionary changes in the frequencies of genes in populations over time? We decided to study the large numbers of Adélie penguins nesting around the Antarctic coastline during summer, and the bones of their ancestors buried beneath these colonies.
At Inexpressible Island (so named because the early explorers could not think of a word bad enough to describe it), our research team took blood samples from a large number of penguins in the colony there. With the help of our collaborator, Professor Carlo Baroni from the University of Piza, we also collected sub-fossil penguin bones and radiocarbon dated some of these to determine their age – estimated to be around 6000 years old.
Next, we isolated nine sets of nuclear DNA sequences from both the modern and the ancient samples and examined the sequences for changes in the frequencies of different gene variants ('alleles'). In fact, we did record some changes in the frequencies of alleles over time. This was the first demonstration of evolution in the sense of gene frequency shifts over a 6000 year period. Other studies have shown changes over much shorter time periods.
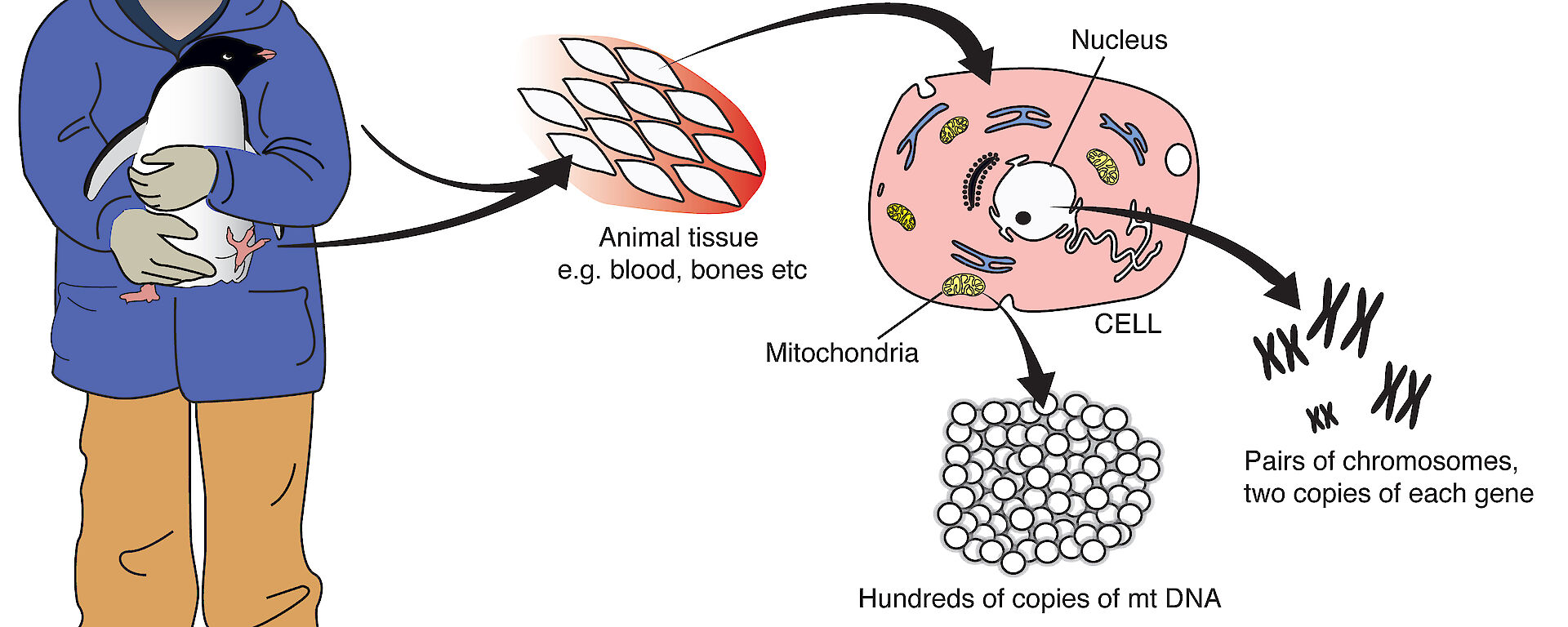
As part of our study we also wanted to measure the speed or rate of evolution; that is, how fast changes in the genetic composition of the penguin population occurs. We took a new approach to the problem by comparing mitochondrial DNA sequences from living birds, with those from sub-fossil bones up to 44 000 years old, excavated from beneath the colonies. (Mitochondria are small structures within cells that provide most of a cell’s energy).
By comparing the modern and ancient mitochondrial DNA sequences, we were able to directly measure changes in DNA lineages over time and estimate the speed of evolution. Surprisingly, we were able to show that the rate of mitochondrial evolution is about six times faster than has been estimated using other methods.
Why might DNA sequence evolution be so fast? One possibility is that Adélie penguins have a very high rate of molecular evolution, simply because they have a high rate of mutation for this part of the mitochondrial genome?
To answer this question, we spent four summers in the Antarctic collecting samples from pairs of breeding penguins. Then, using blood samples from mothers and their chicks, we sequenced the same piece of mitochondrial DNA for which we had measured the speed of evolution using the 44 000 year old sub-fossil bones.
Mitochondrial DNA is inherited quite differently to the nuclear DNA of chromosomes. While offspring inherit nuclear DNA from both parents, they typically inherit mitochondrial DNA only from their mother. So in this experiment, we expected to find the same mitochondrial DNA sequence in all chicks as we found in mother Adélie penguins; except of course for any mutations. As mutations are rare, we did not expect to find many of them.
However, we were surprised to find a large number of mutations. Using new analytical methods developed by Professor Mike Hendy and his colleagues from the Allan Wilson Centre for Molecular Ecology and Evolution in New Zealand, we were able to use this large number of observed mutations to estimate the overall rate of mutation in penguins. This mutation rate turned out to be the same, statistically speaking, as the rate of evolution that we had determined using old penguin bones. Our findings suggest that mutations are the driving force for the high evolutionary rate in Adélie penguins.
These findings suggest that estimates of both mutation and evolution might well be similar in other species and that we can use rates of ‘neutral’ mutation (where there are no differences between the original gene and the mutant) as a proxy for evolutionary rates, when these data are available. Finally, our findings mean that these new molecular techniques can be reliably used to time at least relatively recent divergence events; that is, the genetic separation of related organisms from a common ancestor.
DAVID LAMBERT1 and CRAIG MILLAR2
1Professor of Evolutionary Biology, Griffith University, Nathan, Brisbane, Australia
2 Allan Wilson Centre for Molecular Ecology and Evolution, School of Biological Sciences, University of Auckland, New Zealand
For more information and a list of scientific publications see:
- Professor David Lambert’s publications page
- The University of Auckland School of Biological Sciences